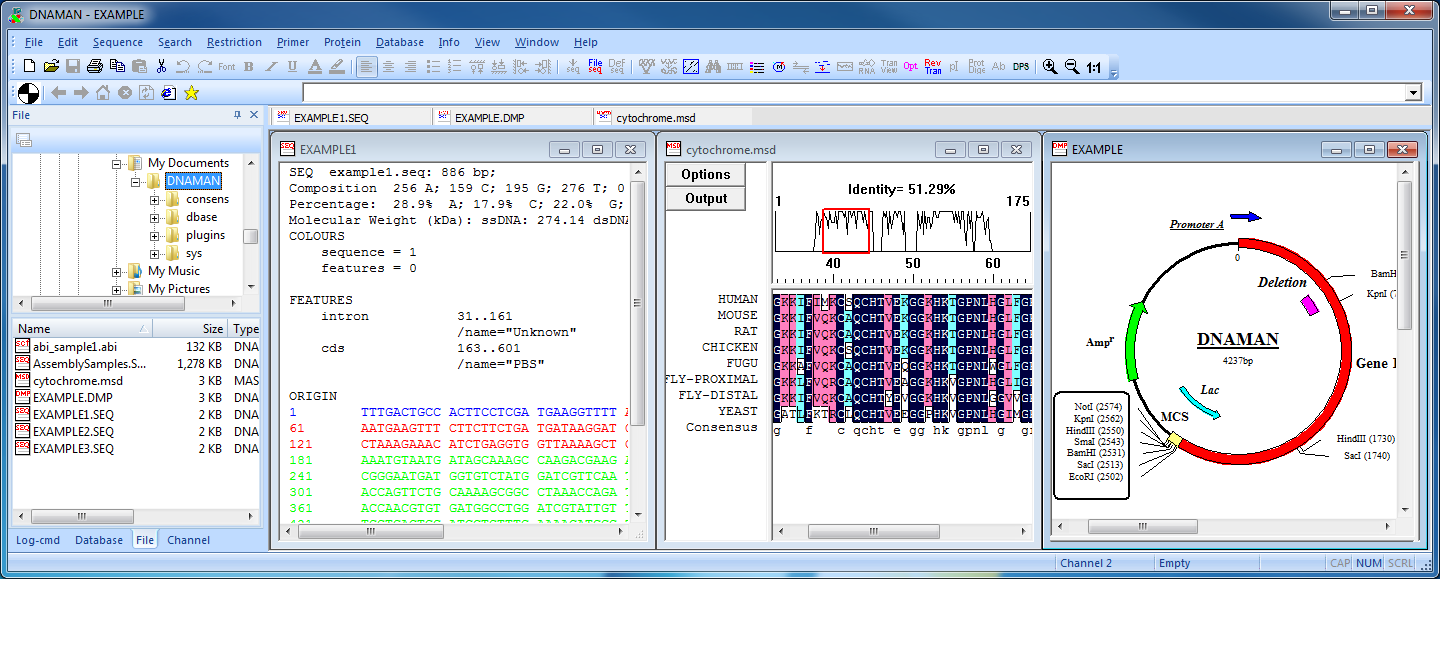
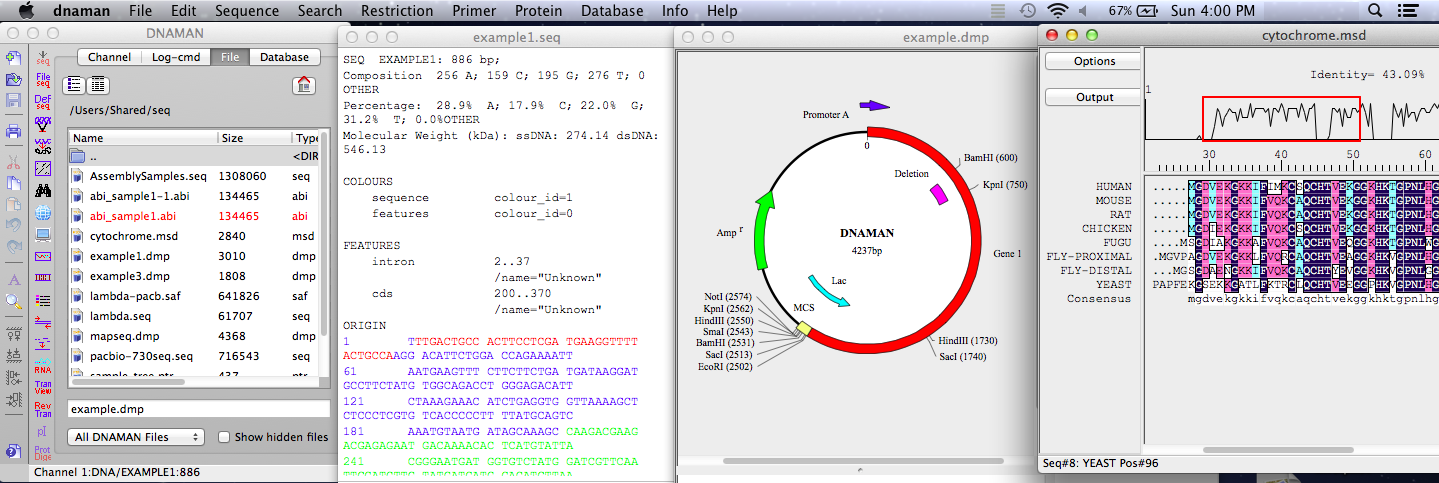
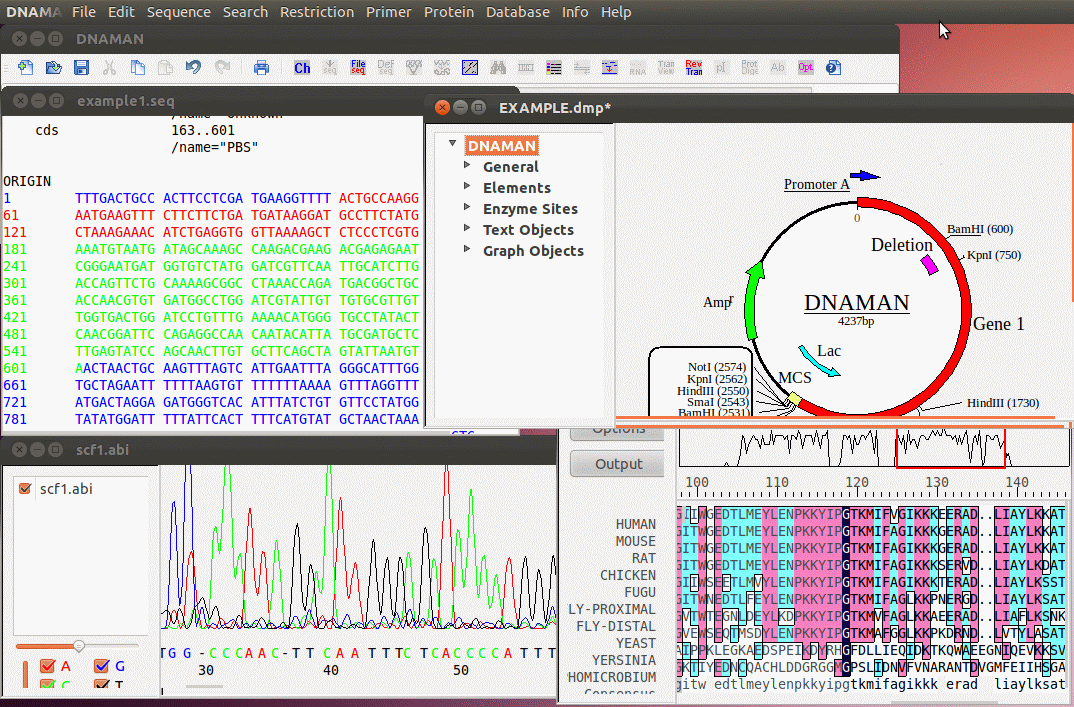
DNAMAN 是一款專門用於開發的高度集成化的分子生物學應用軟體,幾乎可完成所有日常核酸和蛋白質序列分析工作,包含多重序列比對,列如多序列比對、蛋白質分析、限制性酶切分析、質粒繪圖、PCR 引物設計等。這款軟體可便捷的分析各種複雜的運行,還可以直接模擬DNA運行狀態。是生命科學工作者、研究人員所必備的工具之一。支持的分析類型非常豐富,包括蛋白質分析、分子研究、引物分析、序列對比研究等模塊,DNAMAN掃描所有的序列記錄在默認的資料庫搜索當前序列的同源序列。DNAMAN軟體的資料庫功能,允許用戶組織DNA和蛋白質序列的不同科目。DNAMAN常在科學期刊中被高度引用的序列分析軟體。
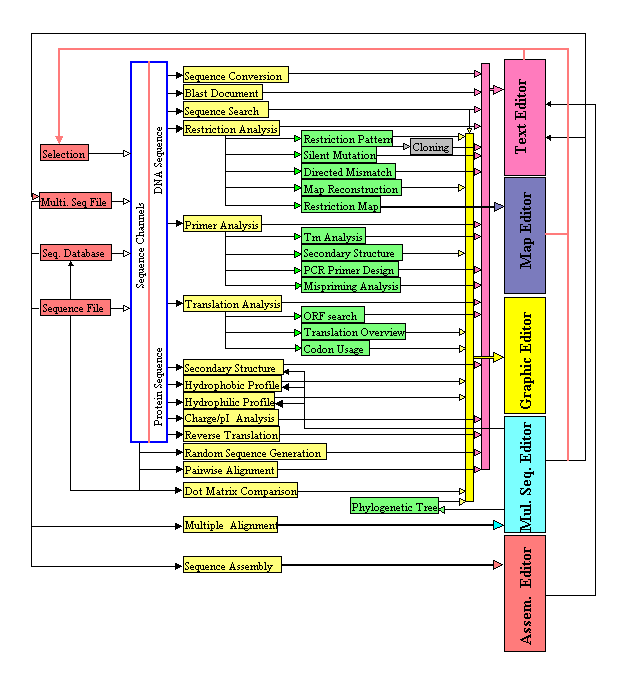
DNAMAN Features
- Integrated Sequence Analysis System
- Sequence Editing
- Restriction Analysis – In Silico Cloning
- Multiple Sequence Alignment
- Phylogenetics Analysis
- Dot-Plot Analysis
- Sequence Assembly
- PCR Primer Selection
- Protein Sequence Analysis
- Database – Sequence Management
Sequence Handling and Conversion
Sequence input
DNAMAN sequences are plain text files formatted with keywords. The program accepts other common sequence formats such as GenBank, GCG, CUSTAL, FASTA, PIR and GDE. DNAMAN utilizes sequence channels to keep active sequences in memory for fast computing. Databases are also available to help organizing sequences for specific research projects. All three sequence input interfaces are accessible on DNAMAN Desktop.
Sequence composition and conversion
DNAMAN reports sequence composition and molecular weight using simple pull-down menu command. It may convert a sequence to its reverse, complementary, reverse complementary, double strand, and RNA sequences.
Sequence Search
DNAMAN provides tools for sequence search and comparison, including nucleotide or amino acid sequences, consensus sequences, open reading frames, repetitive sequences. The small interfering RNAs (siRNAs) selection tool can be used to improve quality of siRNA design. Users can perform BLAST search to NCBI databases or other web based sequence servers within the DNAMAN program.
Restriction Analysis
DNAMAN provide intuitive interfaces for Restriction analysis of DNA sequences. Results can be shown as text, Restriction map and Restriction pattern (Agarose gel). Additional tools include Electronic Cloning, Reconstruction of Restriction maps from fragments, Silent mutation and Directed mismatch.
Sequence Assembly
DNAMAN uses fast and accurate alignment algorithms to assemble a large number of sequences. Trace files can be directly used for assembly. SNPs can be verified through assembled sequence set.
Sequence Homology Analysis
Dot matrix plot
Large sequences can be analyzed in DNAMAN for homologous regions using dotplot. Sequence regions of interest can be zoomed-in and directly aligned.
Two sequence alignment
DNAMAN provides many fast or optimal algorithms to align two DNA or protein sequences.
Multiple Sequence Alignment
DNAMAN uses ClustalW algorithm (Feng-Doolittle and Thompson) for Optimal Alignment, and the global alignment algorithm (Wilbur and Lipman) for fast alignment. The three types of Optimal Alignment in DNAMAN provide high quality alignment results. With the Fast Alignment method, you may quickly align a large number of DNA or protein sequences. Alignment can be further modified or tuned in Multiple Alignment Sequence Editor.
Phylogenetic Analysis
DNAMAN uses common algorithms to calculate the homology matrices and establishes related distances between all pairs of sequences. It can produce distance matrices, and draw phylogenetic trees or homology trees from multiple alignments. Bootstrapping analysis is available for the confidence value of a phylogenetic tree.
Primer Analysis
PCR Primer Design
DNAMAN provides numerous control criteria for PCR primer selection. You may optimize primer filtration by setting the regions of target DNA, size of PCR products, primer characteristics, reaction conditions and primer configurations.
Other Functions
Oligo primers can be analyzed for melting temperature, complementarity, mispriming, silent mutation and directed mismatch.
DNAMAN Database Management
DNAMAN uses standard relational database engine Sqlite3 to manage DNA/Protein sequences and oligo sequences.
| Features | DNAMAN X | DNAMAN XL |
| Sequence Editing and Conversion | Yes | Yes |
| Supporting GenBank, FASTA, GFF and other files | ||
| Sequence conversion with command line and scripting | No | Yes |
| Multiple Sequence Alignment | Yes | Yes |
| Supporting ClustalW, GCG, GDE and other files | ||
| Multiple Alignment with command line and scripting | No | Yes |
| Sequence Assembly | Yes | Yes |
| Fast alignment algorithm with graphic interface | ||
| Sequence Assembly with command line and scripting | No | Yes |
| PCR Primer Design | Yes | Yes |
| Optimized selection with graphic interface | ||
| Batch processing and scripting | No | Yes |
| Sequence Database | Yes | Yes |
| Sqlite3 database engine | ||
| Supporting MySQL and Microsoft SQL server | No | Yes |
| DNA Restriction Analysis | Yes | Yes |
| Intuitive user interface with customizable enzyme database | ||
| Restriction analysis with command line and scripting | No | Yes |
| Pairwise Sequence Analysis and Dot Plot | Yes | Yes |
| DNA/RNA Secondary Structure Analysis | Yes | Yes |
| Sequence Search and BLAST Analysis | Yes | Yes |
| Phylogenetic Analysis | Yes | Yes |
| Protein Sequence Analysis | Yes | Yes |
| Protein Secondary Structure Prediction | Yes | Yes |
| Translation/Reverse Translation/Codon Optimization | Yes | Yes |
| Oligo Sequence Analysis | Yes | Yes |
| Sequence Annotation and Transfer | Yes | Yes |
| Batch Processing and General Scripting | No | Yes |
| Custom Menu Commands | No | Yes |
| Premium Support Services | No | Yes |
| Providing in-depth knowledge with 24-hour response time |